Dataset Description
The Global Ecosystem Dynamics Investigation (GEDI) biomass map is a 1x1 km (“1-km” hereafter) spatial resolution mean aboveground biomass density (AGBD) map derived from GEDI lidar data (Release 2) calibrated using an international network of field inventory plots combined with coincident airborne laser scanning data. These products were developed by the University of Maryland in collaboration with the U.S. Forest Service and the NASA Goddard Space Flight Center.
The GEDI mission characterizes ecosystem structure and dynamics to enable radically improved quantification and understanding of the Earth’s carbon cycle and biodiversity. The GEDI lidar instrument is optimized for mapping canopy structure and aboveground biomass across Earth’s tropical and temperate forests between 51.6° N and 51.6° S latitude. Such maps support a broad range of scientific, policy, and management applications.
A statistical framework allows closed-form estimation of mean AGBD and its uncertainty at the scale of 1-km grid cells, and at coarser scales up to user defined grid cell size or areal extent. GEDI currently provides 1-km estimates of mean AGBD (Mg ha-1) and the uncertainty of these estimates are expressed as the standard error of the mean. The 1-km product does not have estimates for all grid cells because GEDI has not yet achieved global coverage at that spatial resolution. These represent our best understanding of the spatial distribution of Earth’s tropical and temperate forest biomass.
Usage
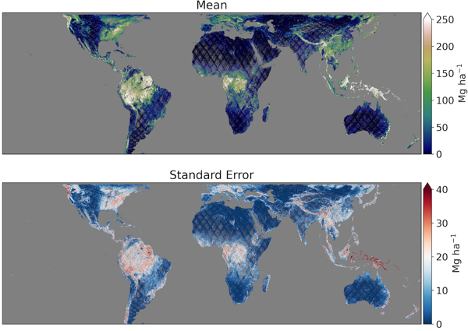
The 1-km dataset consists of two raster files: (a) mean ABGD; and (b) the standard error of the mean, both of which are in units of Mg ha-1. The AGBD estimate is for the entire cell, however some users will require an estimate associated only with the forested part of the cell. Such users may determine what fraction of the cell is forest using the fine-resolution map of their choice and divide the AGBD estimate by that fraction to arrive at a mean for the forested area. By not pre-defining a sub-kilometer forest/non-forest mask, GEDI allows users to choose whichever mask best suits their application. Further details are provided in the GEDI Level-4B (L4B) Algorithm Theoretical Basis Document (ATBD).
For grid cells with valid estimates, the estimated standard error of the mean depends upon the fit of footprint biomass models (model variance) and the density of observations and ground tracks (sample variance). Grid cells without a valid mean biomass estimate have no data (-9999). The distribution of no-data cells is not uniform, with higher non-response found: 1) earlier in the mission life; 2) closer to the equator where the ISS overpass pattern is sparser; 3) in cloudy areas; and 4) in areas where reference ground tracks were not sampled because of an orbital sampling problem experienced in the second year of the missions. This orbital sampling problem involved repeated coverage of some ground tracks at the expense of others because of an unscheduled change in ISS altitude.
Methodology
The GEDI product use the GEDI Level-4A (Release 2) footprint biomass product as input. The GEDI L4A algorithm converts each high-quality surface waveform to an AGBD prediction, and the L4B product uses the sample present within the borders of each 1-km cell to statistically infer mean AGBD. The algorithm used to generate L4A footprint biomass predictions is described in the ATBD for GEDI Level-4A Footprint Aboveground Biomass and the algorithms used to generate the height metrics used as predictors for AGBD are described in the ATBD for GEDI Transmit and Receive Waveform Processing for L1 and L2 Products. ATBD’s are available through the
GEDI website.
A hybrid model-based mode of inference is used in the L4B product. The corresponding 1-km estimates of the standard error of the mean propagate the uncertainty due to both GEDI’s sampling of the 1-km area (as opposed to making wall-to-wall observations) and the fact that L4A biomass values are modeled in a process subject to error instead of measured in a process that may be assumed to be error-free. GEDI ground tracks are treated as cluster samples under the hybrid inference paradigm, and at least two clusters are required to create a valid estimate.
The 1 km2 resolution global EASE-Grid 2.0 is used to partition the GEDI footprint observations into grid cells by their footprint center location. This nested grid features equal-area cells and compatibility with many existing biosphere data sets. More information on this grid can be found from the U.S. National Snow and Ice Data Center (NSIDC) at
https://nsidc.org/data/ease.
Uncertainty and Accuracy
GEDI’s Level-1 (L1) science requirement states that 80% of 1-km cells must be estimated to within a standard error of either 20 Mg/ha or 20% of the estimate, whichever is greater. The Quality Flag variable (2 = meets L1 requirement) facilitates tracking of mission progress with respect to this goal. Just as is the case with designed samples used by the world’s national forest inventories, including the Forest Inventory and Analysis program of the US Forest Service, uncertainty (the standard error of the estimate) is assessed using sample theory in conjunction with observed sample numbers and variances.
Uncertainty of the estimated mean biomass is determined through the estimator described by hybrid estimation. The hybrid variance estimator has two components, the first of which is model variance from the L4A field-to-GEDI AGBD model. The second variance component relates to GEDI’s sample design. These variance components allow the user to decompose uncertainty expressed in the Standard Error variable into its primary components.
Dataset Sustainment
This is a GEDI Release 2 product that will be published at NASA’s ORNL DAAC, with an expected release in October 2021. Annual updates to these products are planned to incorporate new observations collected by GEDI and improvements to the L4A and L4B algorithms. The GEDI mission was recently awarded an extension and is expected to be collecting data until at least January 2023.
Technical Characteristics
Spatial resolution: 1 km
Geographical coverage: 51.6° N and 51.6° S
Temporal coverage: 18-Apr-2019 to 09-Jun-2021
Update frequency: Annual
Format: GeoTIFF
Data Policy: Creative Commons Attribution 4.0 International (CC-BY-4.0)
Associated Guidance or User Manual
User Guide
Data
Points of contact for queries
John Armston
Associate Research Professor
University of Maryland, College Park, USA
Email: armston@umd.edu
Sean Healey
Research Ecologist
U.S. Forest Service
Email: seanhealey@fs.fed.us
Ralph Dubayah
Professor
University of Maryland, College Park, USA
Email: dubayah@umd.edu
